Scaling Down to Go Big: An Overview of the Lab-on-a-Chip
By Julianne Davis - Technical Writer and Support Scientist at Biotium
The Rise of the Lab-on-a-Chip
With the advent of microchips, computers that once spanned whole rooms now fit into the palm of our hand. Cutting-edge science aims to scale down research experiments in a similar manner using microtechnology. Lab-on-a-chip (LOC) technology was born from this goal in 1979 and has evolved and grown in popularity significantly since then.1
LOC is a device that scales single or multiple lab processes down to mm-scale integrated circuits, or chips, to achieve automation and high-throughput screening.2 The development of these devices requires expertise in microfluidics, the study of the physical behavior of minute volumes of fluids. Manipulating reagents at this level allows scientists to exploit microscale effects like rapid heating and mixing, significantly reduce waste, and minimize exposure to dangerous chemicals.1 More recently, this technology has evolved to include micro-engineered single or multi-cell culture platforms. These devices mimic the in vivo environment and have become effective models of living cells and organs.1-2 Consequently, the concept of LOC has gained attention as a powerful tool for biological and biomedical research.3
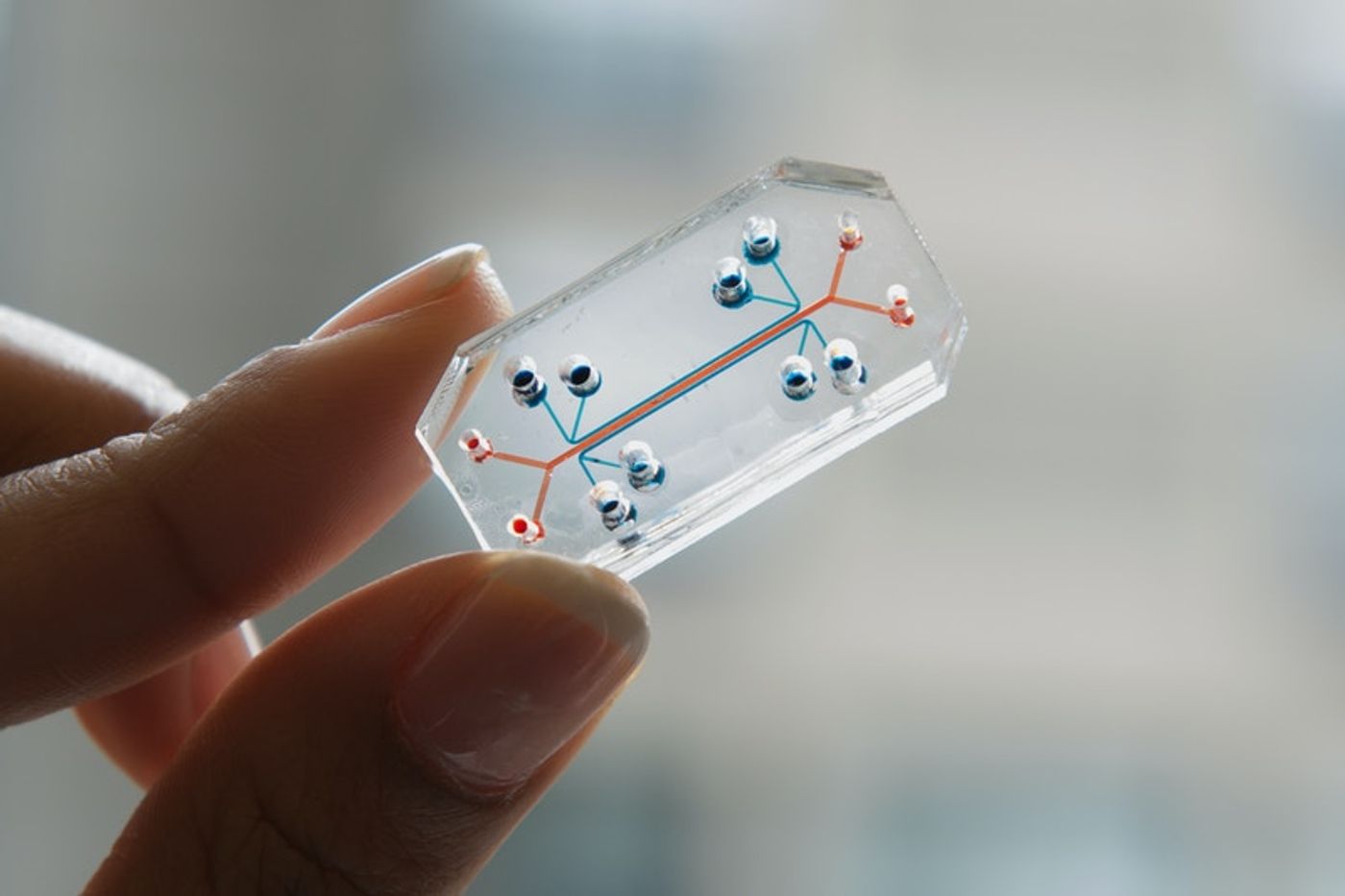
A lung-on-a-chip microdevice. Image courtesy of Wyss Institute/Design Museum.
From Single Synapses to Multiple Organs
Three-dimensional (3D) cell culture technologies are still often used for drug development as these paradigms allow for more biologically relevant cell networks and high-throughput screening.4 However, there are many functional aspects of cells and organs that 3D culture models cannot replicate. These limitations include a lack of tissue-to-tissue interfaces, spatiotemporal gradients of chemicals, and mechanically active components. The application of LOC technology breaks through these barriers.5
Even tricky problems, such as mimicking physically separated biological structures, can now be tackled with LOC. Yamamoto et al. showed this in their use of microdevices to create precisely compartmentalized co-cultures of motor neurons and skeletal muscle fibers to form viable neuromuscular junctions. They showed that these cells are functionally identical to their in vivo counterparts in the presence of an excitatory neurotransmitter, inhibitory chemicals, and mechanical force. These cells can be labeled with fluorescent dyes and imaged on-chip with minimal adjustments to standard protocols, as shown by the use of lipophilic carbocyanine dyes, like CellBrite® Green, to stain motor neuron axons in this system.6
As scientists gained the ability to generate increasingly complex systems on chips, they began creating organoids, leading to the birth of the organ-on-a-chip (OOC) subfield.3 With these stem cell-derived, self-organizing miniature organs, researchers seek to replicate the critical structural and functional characteristics of their in vivo counterparts. Functional OOC could provide cost effective, high-throughput disease model systems as an alternative for drug testing in animals or human tissue.7
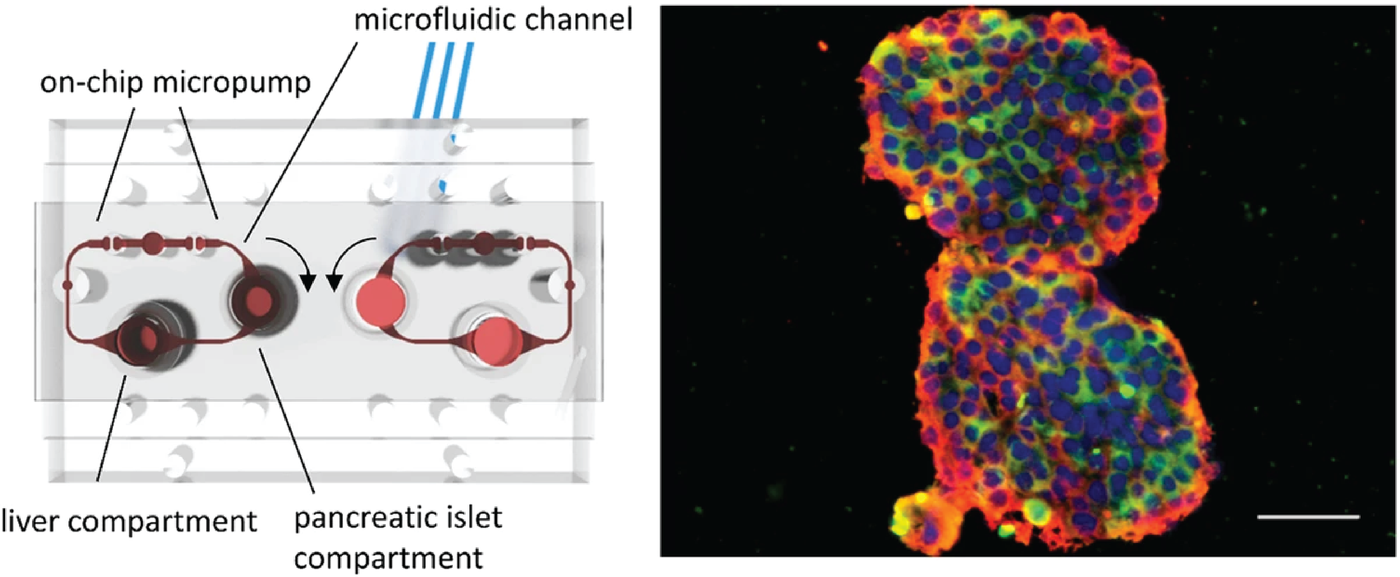
Bauer et al. demonstrated the potential for complex OOC by generating human pancreatic islets and liver spheroids on-chip. This two-organ-chip model, complete with a working feedback loop between the liver spheroids and the insulin-secreting islet microtissues, gave the researchers the ability to study pancreatic islet–liver cross-talk based on insulin and glucose regulation. The robustness and reproducibility was validated by conducting experiments in two independent laboratories, making it a promising type 2 diabetes model.8
An illustration of the two-organ-chip used by Bauer, et al. with media circuits, respective culture compartments, and micropump valves highlighted in red. The panel on the right shows the pancreatic islet microtissues and liver spheroids used in the experiment immunostained for insulin (pseudocolored red) and glucagon (CF®594, pseudocolored green). Credit: Bauer, et al. https://doi.org/10.1038/s41598-017-14815-w reproduced under the Creative Commons license.
The Future of LOC Technology
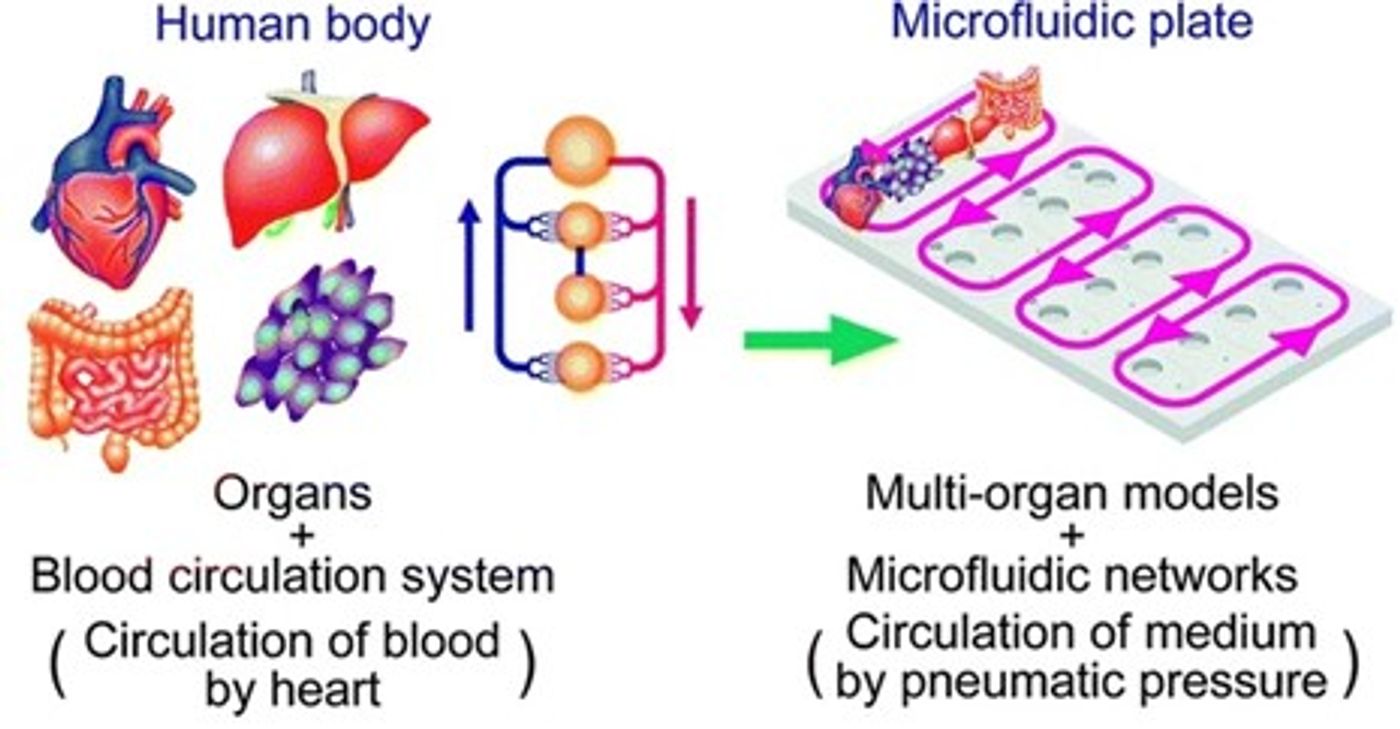
While LOC and OOC have laid the groundwork for many potential methods of revolutionizing medicine and scientific research, most current end goals still lie on the horizon. Multi-organ chips are currently at the cutting edge of this field. To fully reflect the complexity, functionality, and integrity of an organ, one must account for the array of physiological pathways and inter-tissue interactions that exist in vivo.1 With multi-organ chips, like the liver and pancreatic circuit demonstrated by Bauer et al., researchers seek to simulate physically interconnected multi-organ systems.8
A representative multi-throughput multi-organ chip system. Credit: Wu, et al. https://doi.org/10.1186/s12938-020-0752-0 reproduced under the Creative Commons license.
Experts hope that multi-organ chips will eventually lead to the creation of patient-on-a-chip technology. This precise approach would entail creating a complex, individualized multi-organ chip model allowing clinicians to develop treatment plans tailored to each patient’s unique biology. It would also create an attractive alternative to the use of animal models. While many limiting factors still exist, including the expense of the components, the need for universal cell culture media, and the challenge of managing and interpreting data from such a complex system, the potential reward of revolutionizing our approach to personalized medicine and laboratory research keep scientists pushing forward.1
References
- Wu, Q., Liu, J., Wang, X., Feng, L., Wu, J., Zhu, X., Wen, W., & Gong, X. (2020). Organ-on-a-chip: recent breakthroughs and future prospects. Biomedical Engineering Online, 19(1), 9. https://doi.org/10.1186/s12938-020-0752-0
- Volpatti, L. R., & Yetisen, A. K. (2014). Commercialization of microfluidic devices. Trends in Biotechnology, 32(7), 347–350. https://doi.org/10.1016/j.tibtech.2014.04.010
- Azizipour, N., Avazpour, R., Rosenzweig, D. H., Sawan, M., & Ajji, A. (2020). Evolution of Biochip Technology: A Review from Lab-on-a-Chip to Organ-on-a-Chip. Micromachines, 11(6), 599. https://doi.org/10.3390/mi11060599
- Langhans S. A. (2018). Three-Dimensional in Vitro Cell Culture Models in Drug Discovery and Drug Repositioning. Frontiers in Pharmacology, 9, 6. https://doi.org/10.3389/fphar.2018.00006
- Huh, D., Hamilton, G. A., & Ingber, D. E. (2011). From 3D cell culture to organs-on-chips. Trends in Cell Biology, 21(12), 745–754. https://doi.org/10.1016/j.tcb.2011.09.005
- Yamamoto, K., Yamaoka, N., Imaizumi, Y., Nagashima, T., Furutani, T., Ito, T., Okada, Y., Honda, H., & Shimizu, K. (2021). Development of a human neuromuscular tissue-on-a-chip model on a 24-well-plate-format compartmentalized microfluidic device [10.1039/D1LC00048A]. Lab on a Chip, 21(10), 1897-1907. https://doi.org/10.1039/D1LC00048A
- Park, S. E., Georgescu, A., & Huh, D. (2019). Organoids-on-a-chip. Science (New York, N.Y.), 364(6444), 960–965. https://doi.org/10.1126/science.aaw7894
- Bauer, S., Wennberg Huldt, C., Kanebratt, K. P., Durieux, I., Gunne, D., Andersson, S., Ewart, L., Haynes, W. G., Maschmeyer, I., Winter, A., Ämmälä, C., Marx, U., & Andersson, T. B. (2018). Publisher Correction: Functional coupling of human pancreatic islets and liver spheroids on-a-chip: Towards a novel human ex vivo type 2 diabetes model. Scientific Reports, 8(1), 1672. https://doi.org/10.1038/s41598-017-14815-w