Reduce the risk of mycoplasma contamination in cell cultures
Introduction
Mycoplasma contamination is prevalent in cell culture labs: in one study, researchers identified mycoplasma contamination in 11% of the 10,000 cell lines they tested (Olarerin-George et al. 2015). Part of the reason mycoplasma contamination is so prevalent is that mycoplasma spreads efficiently even in routine subculturing. In another study, researchers detected live mycoplasma outside of the flask, on a hemocytometer, on the pipette, and outside of the pipette discard pan after working with mycoplasma-infected cell culture in a laminar flow hood. Indeed, live mycoplasma were recovered from the surface of the laminar flow hood even four to six days later. And a clean culture that was subcultured once a week in the same hood following work with the contaminated cells tested positive for mycoplasma after only 6 weeks (reviewed in Nikfarjam et al 2012, original article McGarrity GJ (1976).
What are mycoplasma?
Mycoplasma are a genus of bacteria that lack a cell wall. Because they lack a cell wall, they can take on different shapes, from round to elongated, and they are resistant to antibiotics (Bebear et al. 2011). Due to their small size (0.2 – 0.8 μm), mycoplasma can penetrate sterilizing-grade filters, making them a very difficult contaminant to eliminate. Their small size also makes them difficult to spot within cell cultures (Masters et al. 2007; Volokhov et al. 2008).
In cell culture, mycoplasma inhibit protein biosynthesis and cell growth and alter RNA and DNA synthesis, thus they can affect research results. In cell-based therapeutics and in advanced therapy medical products (ATMPs), mycoplasma contamination can cause immune reactions, chromosomal aberrations, or altered proliferation characteristics, making therapy unsafe for use in patients (Martins et al. 2014).
Potential routes for mycoplasma contamination include:
- Contaminated media, reagents or instruments
- Laboratory personnel (approximately 80.6% of technicians are mycoplasma carriers)
- Infected cultures obtained from other labs
Best practices to avoid mycoplasma contamination
You can achieve contamination-free cell cultivation with these practical procedures:
1. Implement Good Cell Culture Practice
Good Cell Culture Practice (Coecke S, 2005) is a set of general cell culture guidelines to prevent contamination without needing to resort to the use of antibiotics. Routine use of antibiotics in cell culture media is not recommended. Not only are antibiotics often ineffective against mycoplasma, but they also promote antibiotic resistance and can hide low-level persistent contamination (Drexler & Uphoff, 2002). In addition, antibiotics in media may have an effect on the outcome of cell culture experiments.
Key guidelines in Good Cell Culture Practice include: cultivation of only one cell line at a time; clearly marking the pipettes, pipetting controllers, and pipette tips used for cell culture work; only using these pipettes and tips in the cell culture laboratory; and never moving them from the cell culture laboratory to another and back again. Only use reagents designated exclusively for cell culture work, and do not use stock solutions designated for other applications.
2. Select easy-to-clean and autoclavable pipettes for cell culture work
Maintenance of pipettes and liquid handling controllers play an important part in reducing risks of spilling and splashing of liquids, a key source of contamination. Laboratories should set guidelines and protocols for regular decontamination and maintenance. According to Eterpi et al. (2010), effective cleaning methods for removing bacterial contaminations are:
- Autoclaving
- Alkaline cleaners
- Ethanol Vaporized hydrogen peroxide VHP
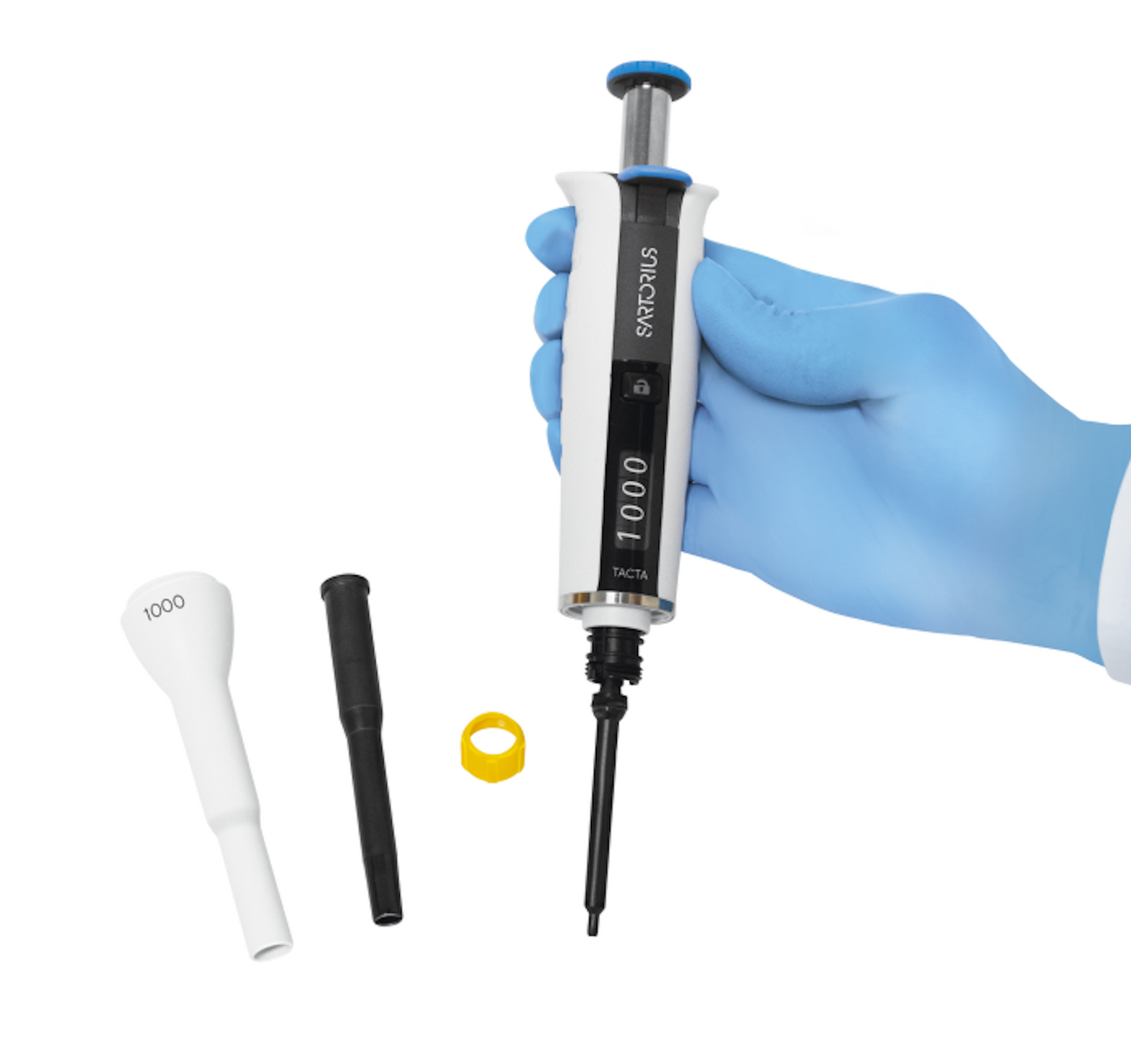
There are significant differences between pipette designs that affect their cleaning protocols. For instance, depending on the design, a disassembled pipette could be between three or more than twenty parts. To make cleaning and sterilizing easy and reliable, choose pipetting tools that are fully autoclavable, easy-to-clean, and tolerate effective cleaning methods, such as the Sartorius Tacta® pipette.
Filter tips are the safest choice for cell culture work, as they provide the best protection for both the samples and the pipette, and they prevent mycoplasma spreading by aerosols. Autoclaving is not suitable for filter tips because the filter material, polyethylene, does not tolerate high temperatures. Therefore, pre-sterilized pipette tips are most commonly sterilized with electron beam (beta) irradiation. Sartorius uses e-beam pre-sterilization with a sterility assurance level (SAL) of 10-6. This means that there is a maximum probability of one viable microorganism in 1 million sterilized items. This SAL level is accepted for pharmacopoeial sterilization procedures. Sartorius evaluates the e-beam radiation dose four times a year with independent dose audits.
Dust and dirt collect on the surface of tip boxes when storing tip racks in warehouses and lab shelves. Therefore, choose individually wrapped tip boxes, because the wrapping ensures sterility during storage. Only open the wrapping just before use and put the unwrapped box straight into the laminar flow cabinet to minimize carryover of dirt to the clean working area.
4. Test regularly
Test all cell lines regularly for mycoplasma contamination, especially new cell lines. Keep new cell lines isolated in a separate incubator until they can be reliably shown free of mycoplasma. Many researchers prefer using mycoplasma PCR/qPCR kits because they are very sensitive, and results can be obtained fast. As PCR/qPCR cyclers are common in most laboratories, the testing is easy to conduct.
Request a free trial of our Microsart® ATMP Mycoplasma kit here: https://www.sartorius.com/en/applications/quality-control-testing/microbiological-quality-control/rapid-microbial-testing#id-1177396
5. Filter with 0.1 µm filter
Autoclaving is an efficient way to kill mycoplasma, but unsuitable for some media and reagents, because heat destroys many nutrients and growth factors. Standard sterile filtration with pore size of 0.22-μm that is generally efficient in preventing the passage of bacteria and fungi is inadequate for mycoplasma that have a cell size of 0.2-0.8 μm and can pass through pores of > 0.1 μm in size. Therefore, to prevent mycoplasma contamination, use the 0.1 μm pore size instead of standard 0.22 μm sterile filters when filtering cell culture reagents and media (Roche et al. 1992).
References
Bebear C, Pereyre S, Peuchant O. Mycoplasma pneumoniae: susceptibility and resistance to antibiotics. Future Microbiol (2011); 6; 423–431.
Coecke S, Balls M, Bowe G, Davis J, Gstraunthaler G, Hartung T, Hay R, Merten O-W. Guidance on Good Cell Culture Practice. ATLA (2005); 33; 261-287.
Drexler HG & Uphoff CC. Mycoplasma contamination of cell cultures: Incidence, sources, effects, detection, elimination, prevention. Cytotechnology (2002); 39; 75-90.
Eterpi M, McDonnell G, Thomas V. Decontamination efficacy against Mycoplasma. Letters in Applied Microbiology (2010); 52; 150-155.
Martins JP, Santos JM, de Almeida JM, Filipe MA, de Almeida MV, Almeida SC, Água-Doce A, Varela A (2014). Towards an advanced therapy medicinal product based on mesenchymal stromal cells isolated from the umbilical cord tissue: quality and safety data. Stem Cell Res Ther (2014); 5(1); 9.
Masters JR, Stacey GN. Changing medium and passaging cell lines. Nat. Protoc. (2007); 2; 2276–2284.
McGarrity GJ. Spread and control of mycoplasmal infection of cell cultures. In vitro (1976); 12 (9); 643-648.
Nikfarjam L & Farzaneh P. Prevention and Detection of Mycoplasma Contamination in Cell Culture. Cell J (2012); 13(4); 203-212.
Olarerin-George AO & Hogenesch JB. Assessing the prevalence of mycoplasma contamination in cell culture via a survey of NCBI’s RNA-seq archive. Nucl Acid Res (2015); 43(5); 2535-2542.
Roche KL, Levy RV. Methods used to validate microporous membranes for the removal of Mycoplasma. Biopharm 1992:22-33.
Volokhov DV, Kong H, George J, Anderson C, Chizhikov VE. Biological enrichment of mycoplasma agents by cocultivation with permissive cell cultures. Appl. Environ. Microbiol. (2008); 74; 5383–5391.